What benchmarks miss: two clinical AI failures that reshaped how we build medical LLMs
*Originally published November 2023 in Medhealth Outlook.*
Passing the US medical licensing exam got generative AI its healthcare headline. Running in a hospital’s information-extraction pipeline is a different story. By late 2023, peer-reviewed evaluations on clinical named entity recognition, social determinants extraction, and de-identification were landing one after another, each with the same finding: GPT-4 trailed task-specific models built for the job. Two cases we worked on pushed that finding from benchmark curiosity to architectural decision. One was adverse-event extraction from opioid progress notes for an FDA Sentinel program. The other was reasoning over a patient’s timeline to decide whether a drug had actually caused a reaction. Both taught us things no leaderboard number would have.
What the medical exam score hides
The USMLE result did real work for the field. It forced clinicians and health-IT buyers to take generative AI seriously, and it seeded the first wave of hospital pilots. The problem is how that result traveled. “GPT-4 passes the medical exam” became shorthand for “GPT-4 is ready for clinical text,” and those two claims have almost nothing to do with each other.
The USMLE is a closed-book, multiple-choice test. Each question is a clean vignette with five answer choices, designed by clinicians to have one defensible right answer. A real discharge note is none of those things. It is between five and twenty pages of mixed narrative and templated text, written by three or four authors over a two-week stay, with copy-and-paste duplication, undocumented abbreviations, implicit negations (”ruled out PE, started DVT prophylaxis anyway”), and section headers that shift meaning depending on where they appear. Production healthcare AI is graded on what the system does with that mess, not on what it does with a board-exam stem.
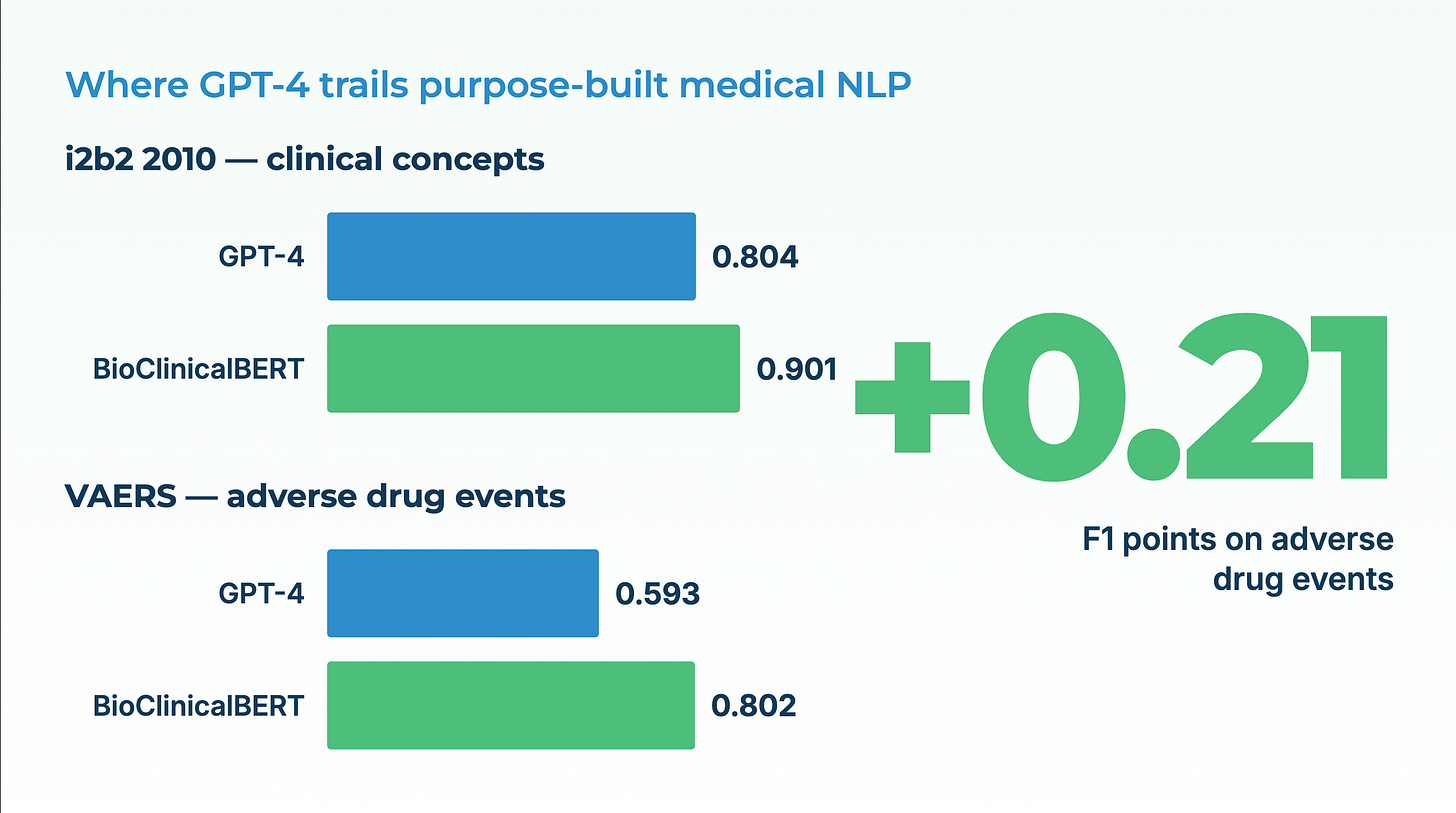
A systematic review published in *Health Care Science* in 2023 catalogs the gap. General-purpose LLMs show competitive performance on benchmark question-answering and summarization tasks, and considerably weaker performance on the entity-level extraction work that actually powers clinical pipelines. A 2024 *JAMIA* study quantified the same pattern on the 2010 i2b2 concept-extraction benchmark: GPT-4 with baseline prompting reached an F1 of 0.804 on MTSamples. BioClinicalBERT, a 110-million-parameter domain-specific model released years earlier, reached 0.901 on the same dataset. On the VAERS adverse-event corpus, the gap was wider: GPT-4 at 0.593 versus BioClinicalBERT at 0.802. Careful prompt engineering closed part of the gap but never eliminated it — and the prompt engineering itself required enough clinical labeling to have trained the domain model in the first place.
The headline-to-reality distance matters because every procurement conversation in healthcare AI starts with it. Executives see the exam number. Engineers inherit the delta.
Lesson one: unstructured notes are where adverse events hide
The first lesson came from a Sentinel Innovation Center program focused on opioid-related adverse events. Sentinel is the FDA’s post-market drug-safety system, and it runs primarily on structured claims data: billing codes, dispensing records, diagnosis flags. For many safety signals, claims are enough. For opioids, they are not.
A clinician who sees a patient nodding off in the chair, documents “appears sedated, spouse concerned about dosing,” and adjusts the prescription downward has just recorded an adverse event. No billing code captures that. The note does. Extracting it requires three linked NLP tasks done together: event classification (is “sedated” describing an observed state, a risk factor, or a ruled-out condition?), named entity recognition (which medication, which dose, which side effect?), and relation extraction (is the sedation linked to the opioid, or to the benzodiazepine started last week, or to neither?).
This is where the general-versus-specialized split stopped being theoretical. On the clinical NER benchmarks that matter for this work, i2b2 2010 for concepts, n2c2 for medications and adverse drug events, domain-specific models held a steady 5 to 15 F1-point lead over general-purpose LLMs, even after prompt tuning. A 2024 *Bioinformatics* paper, BioNER-LLaMA, showed that fine-tuning a 7-billion-parameter open model on biomedical NER data yielded F1 improvements of **5 to 30 points** over few-shot GPT-4 across three standard datasets. The authors noted what anyone who has built these systems knows: LLMs are strong generative models and weaker sequence labelers, because NER is fundamentally a span-localization problem that generation-first models solve awkwardly.
Three observations from our Sentinel work sharpened the point:
The first is that negation and uncertainty dominate clinical text. In a progress note, “denies chest pain” and “c/o chest pain” are one word apart and medically opposite. A general LLM reading under a generation objective tends to smooth over these distinctions, because smooth text is what it was rewarded for producing. A task-specific assertion-status classifier is explicitly trained to tag “chest pain” as *present*, *absent*, *possible*, *conditional*, or *family history*, and the downstream pipeline treats each differently.
The second is that clinical sub-specialties are effectively different languages. A psychiatry progress note and an oncology consult share maybe 40% of their vocabulary. “RA” means rheumatoid arthritis to a rheumatologist and right atrium to a cardiologist; “MS” is multiple sclerosis in neurology and mitral stenosis in cardiology. A model that handles both correctly is one that has seen both, in quantity, with labeled context. Frontier LLMs have seen both; they have not been rewarded for distinguishing them under uncertainty.
The third is that throughput and cost are features. Sentinel work involves hundreds of thousands of notes per study. A 110M-parameter specialized NER model runs on a single commodity GPU at thousands of notes per minute. A frontier LLM running the same task through an API costs orders of magnitude more per note and introduces a per-request latency that turns a three-hour job into a three-day one. For programs with statutory deadlines and bounded compute budgets, that difference decides whether the analysis happens at all.
Lesson two: a list of findings is not an answer
The second lesson came from a different kind of failure. A pharma-safety team wanted to know, for a specific cohort of asthma patients on montelukast, whether neuropsychiatric symptoms documented in the clinical record appeared *after* the drug was started and plausibly followed from it. The information was in the notes. The challenge was not finding it — it was reasoning over the timeline.
Modern extraction pipelines are excellent at pulling structured findings from unstructured text: medication start dates, diagnosis dates, symptom onset, dosage changes. What they do not do out of the box is answer questions that require ordering those findings and reasoning over the order. “Did the insomnia start within 14 days of starting montelukast, and was there a documented attempt to rule out other causes?” is a question no single extraction returns. It requires the system to assemble a per-patient timeline and reason over it.
This is the emerging pattern in medical AI, and it is where LLMs genuinely help — once the underlying data is clean. The architecture that worked for us was a three-layer stack. Small, accurate, task-specific models at the bottom: entity recognition, relation extraction, assertion status, entity resolution to SNOMED and RxNorm. A timeline-assembly layer in the middle: linking the extracted facts into a per-patient longitudinal record, normalizing dates, resolving coreference across notes. And a reasoning layer on top, which can be an LLM, used for what generative models are actually good at, reading an already-structured timeline and answering natural-language questions about it.
When we tried to collapse the stack by sending raw notes plus a reasoning question directly to a general-purpose LLM, accuracy fell. Not because the LLM couldn’t reason, but because the extraction errors it made at the bottom compounded into the reasoning errors it made at the top. A misclassified assertion status on page 3 became a phantom adverse event on page 14. The fix was not a better prompt. The fix was to stop asking one model to do both jobs.
The broader pattern shows up across the peer-reviewed literature. For structured extraction, NER, relation extraction, entity resolution to ICD-10 and SNOMED, domain-specific models win. For summarization, question-answering over already-structured data, and conversational reasoning, LLMs win, especially when the input they operate on has been cleaned up by the specialized layer beneath them. On social determinants of health extraction, a 2024 benchmark found that GPT-4 made roughly three times as many errors as fine-tuned models, because SDOH is a long tail of subtle social context that benefits disproportionately from domain training. On de-identification of clinical notes, domain-tuned models routinely reach above 99% PHI detection while general LLMs sit in the low 90s, with the downstream consequence that one system can run without human review and the other cannot.
What this means for how you build
For executive buyers, CIO, CMIO, CAIO, CDO, this translates into four things worth pushing on when a vendor pitches a medical LLM.
First, ask what benchmarks the accuracy claims come from, and whether those benchmarks include entity-level extraction on clinical text, not just multiple-choice question answering. The headline “passes the medical exam” tells you almost nothing about production performance.
Second, ask for per-task accuracy, not aggregate accuracy. A single number hides the places where general LLMs are genuinely strong (summarization, patient-facing Q&A over a curated knowledge base) and the places where they are not yet strong enough for unsupervised production use (NER, assertion status, de-identification at scale).
Third, ask about the architecture. A system that layers task-specific models for extraction under a generative model for reasoning is a more honest answer than a system that claims one frontier model will do everything. The first will be measurable and improvable per component. The second will be expensive to run and hard to diagnose when it’s wrong.
Fourth, ask about cost and deployment. Running frontier LLMs on every clinical note in a real safety or cohorting program is not economically viable today for most organizations, and in many cases the data cannot leave the environment at all. Specialized models that run on commodity hardware, inside the hospital firewall, are not a second-best option — for production-grade healthcare text work, they are usually the only option that finishes.
The short version
Exam scores made healthcare LLMs credible. Production work made them specific. The two lessons from the field, that unstructured notes are where safety signals actually live, and that a list of extracted findings is not yet an answer, both point the same direction. Purpose-built medical language models, composed into a clean stack, outperform general-purpose LLMs on the clinical tasks that matter and cost a fraction of the price. The work of the next several years is continuing to build that stack, component by measurable component, and resisting the temptation to let a board-exam headline substitute for it.
FAQ
Why does GPT-4 underperform specialized models on clinical NER if it’s the more capable model overall?
Because named entity recognition is a sequence-labeling task and LLMs are trained as text generators. The objective mismatch shows up as over-confident labeling of non-entity spans and under-recall on entities that don’t look like the training distribution. Specialized encoder models (PubMedBERT, BioClinicalBERT) are trained with a labeling objective and domain data. On the 2010 i2b2 benchmark and the VAERS dataset, BioClinicalBERT outperformed GPT-4 by roughly 5 to 15 F1 points depending on the corpus.
Does prompt engineering close the gap?
Partially, not fully. A 2024 *JAMIA* paper showed GPT-4’s F1 on MTSamples rising from 0.804 to 0.861 with a four-component task-specific prompt framework. BioClinicalBERT, without prompt work, sat at 0.901. And the prompt engineering required enough labeled clinical data to have trained a specialized model in the first place.
Where do LLMs earn their place in clinical pipelines?
On summarization of already-structured inputs, conversational question-answering over curated knowledge, and reasoning over an assembled patient timeline. In each case, the LLM operates on clean input produced by specialized models upstream, not on raw clinical text.
Why is de-identification treated as a production bottleneck for general LLMs?
Because the accuracy bar is unusually high and the volumes are unusually large. Below about 99% PHI recall, every note needs human review, which defeats the automation. Above 99%, the pipeline can run unsupervised. General LLMs have not consistently crossed that threshold on real clinical text; domain-tuned systems have, which is what makes hospital-scale de-identification economically viable.
What about the data-privacy side?
Sending clinical notes to a third-party cloud API raises HIPAA, GDPR, and data-sovereignty issues that in many organizations rule the option out before accuracy even enters the conversation. On-premises or private-cloud deployment, with no data leaving the customer’s control, is a hard requirement for most regulated buyers, and most real healthcare AI workloads end up running that way regardless of which model family delivers the best benchmark number.
---
David Talby is CEO of John Snow Labs, whose healthcare NLP and medical LLMs are used by 500+ healthcare and life sciences organizations, including collaborations with the FDA on post-market drug safety. He also leads Pacific AI, which focuses on governance for healthcare AI.